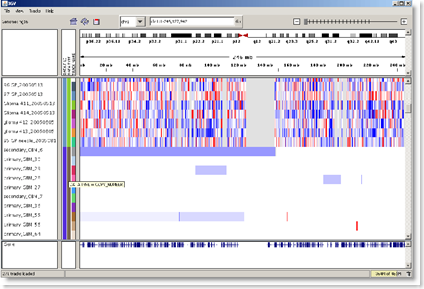
The software offers both detailed, localized views of genomic data, and a whole genome view, in a format somewhat similar to the way Google presents items on the web. It is free to download and use. There are also some sample data files available. I like the idea of the heat map view being available. This should make the analysis process move along a little faster for researchers. BD
NEW YORK (GenomeWeb News) The Broad Institute of MIT and Harvard has created a genomics informatics tool that will allow researchers to visualize genomic information, and has made it publicly available for free, Broad said today. The Integrative Genomics Viewer was developed to help researchers find effective ways to visualize the vast amounts of diverse genomic data that scientists are accumulating with ever increasing speed, according to the Institute. It can incorporate multiple types of genomic data and allows researchers to view them at various levels of resolution.
It can visualize layers of information about gene expression, sequence alterations, mutations, and other genomic details such as copy number variation, chromatin immunoprecipation data, and epigenetic modifications.
The Integrative Genomics Viewer (IGV) is a fast, flexible viewer for genomic data. IGV visually integrates datasets from various platforms and sources, including:
- Genetic variation data (copy number, loss of heterozygosity, somatic mutations)
- Gene/microRNA expression data
- Epigenetic data
- RNAi screens
- Genomic features
- Clinical and phenotype annotation of samples
GenomeWeb News: Broad Institute Makes Genomics Data Viewer Public



Hi--
ReplyDeleteUCSC (Go Banana Slugs!) has offered similar functionality for years--at least of ACTG sequence data and SNP data, if not the RNAi stuff. Here's an example:
http://genome.ucsc.edu/cgi-bin/hgTracks?hgsid=110939747&clade=vertebrate&org=Human&db=hg18&position=chrX%3A151%2C073%2C054-151%2C383%2C976&pix=620&Submit=submit&hgsid=110939747
Here is the main site:
http://genome.ucsc.edu/cgi-bin/hgGateway
I'm not sure how this small "hippie" college got to be a world leader in genomics research, but they have had excellent tools since at least 2003.
In my past life, I was a bona-fide genomics researcher!